Here you see a short segment of an AdhE spirosome from Escherichia coli in the extended, active filament conformation [38].
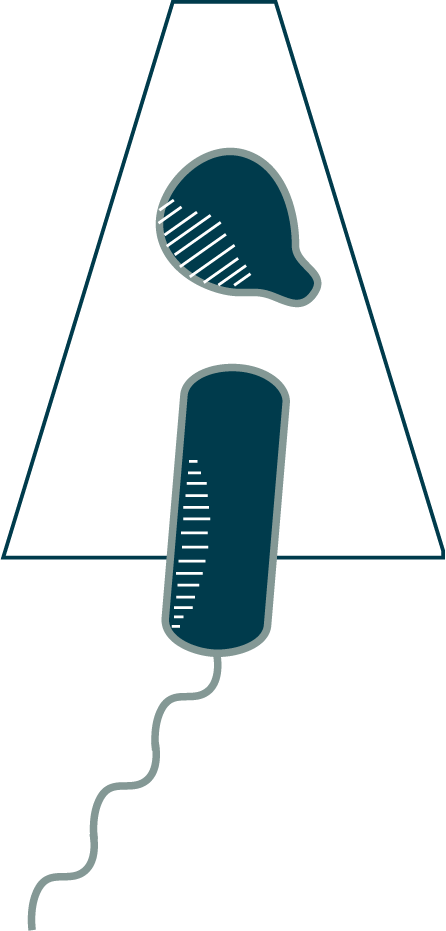

Cells are subject to the vagaries of an environment in which conditions often change. To respond, they must be metabolically agile. The production of enzymes is dialed up and down to meet demand, but they can also be regulated in other ways. Degrading and resynthesizing proteins wastes resources and time. So why not just grant workers a sabbatical when demand lags and call them back in when things pick up again? Cells across all domains of life have evolved an elegant sabbatical mechanism: enzyme polymerization. You already saw an example of this in Chapter 3: the metabolic enzyme CTP synthase. Another example is the alcohol-acetaldehyde dehydrogenase (AdhE) enzyme that allows Clostridium thermocellum like this one to digest cellulose into ethanol. As you can see, AdhE polymerizes into helical filaments called spirosomes. Polymerization can either inactivate an enzyme (e.g. by occluding its active site), or activate it (e.g. by changing its conformation to open an active site). Filaments can also regulate activity by changing their conformation; for AdhE, it is thought that spirosomes torque between a compact, inactive form to a more extended, active form (what you see here) (⇩). Whatever the details, polymerization is a rapid way to mobilize (or demobilize) a large number of enzymes for a metabolic task.
Unoccupied workers can also be recruited to other projects. Remember that CTP synthase filaments play a secondary, cytoskeletal role. (And perhaps many, if not all, cytoskeletal elements evolved this way.) AdhE spirosomes seem to have a secondary function in cell adhesion, although the details are not yet clear.
Here you see a short segment of an AdhE spirosome from Escherichia coli in the extended, active filament conformation [38].